Introduction
In December 2019, a novel coronavirus (SARS-CoV-2) that causes coronavirus disease 2019 (COVID-19) emerged from its epicenter in Wuhan Province in China (1). The rapid spread of this disease seriously threatened global public health and it had a huge impact on the global economy. In the wake of the rapid spread of the coronavirus, there was an urgent need for rapid and sensitive detection systems. Hence, suspected cases were effectively identified, patients were rapidly screened, and virus surveillance was conducted for isolation strategies (2).
The standard method to detect SARS-CoV-2 has been real-time polymerase chain reaction (RT-PCR) (3, 4). Regardless of its high sensitivity and specificity, this method has some drawbacks as it requires complex and expensive equipment, extensive user training, and multiple hours to obtain the result. These weaknesses limit the screening capacity of the RT-PCR technique, and it falls behind rapidly growing SARS-CoV-2 cases (5, 6). As a result, more efficient methods are in high demand for detecting COVID-19 to catch its growing pace.
Recently, reverse transcription loop-mediated isothermal amplification (RT-LAMP) has been used as an alternative to the RT-PCR method. Reverse transcription loop-mediated isothermal amplification has many advantages over RT-PCR, such as requiring a simple instrument (e.g., heating block) and a constant temperature for amplification, giving results in a short time (7). Moreover, the results can be visualized by the naked eye with the presence of a colorimetric pH indicator. Since having the same sensitivity and specificity as RT-PCR, the RT-LAMP method is a more effective choice for high-throughput and low-cost detection of SARS-CoV-2 (8, 9).
This study aimed to compare the efficiencies of RT-PCR and colorimetric RT-LAMP assays to determine if RT-LAMP is suitable for rapid on-site diagnosis of SaRS-CoV-2 based on N-gene.
Materials and Methods
Clinical Sample Handling
Samples were collected as nasopharyngeal swabs in a virus inactivation medium (vNAT; Bioeksen R&D Technologies Ltd, Turkey). vNAT buffer extracted and preserved viral nucleic acids in respiratory tract samples. The sample collection occurred as part of the routine operation of Hacettepe University Faculty of Medicine, and each sample was tested in both RT-LAMP and RT-PCR.
RT-PCR
RT-PCR was performed by using Bio-Speedy SARS-CoV-2 RT-PCR kit (Bioeksen R&D Technologies Ltd, Turkey). Detection with the kit was achieved via rapid nucleic acid extraction from respiratory tract samples followed by multiplex RT-PCR targeting the SARS-CoV-2 specific ORF1ab gene and the human RNase P gene and mRNA in CFX96 real-time PCR instrument (Bio-Rad Laboratories Inc., USA). The oligonucleotide set targeting the human RNase P gene and mRNA functioned as a control of the sampling, nucleic acid extraction, and inhibition.
RT-LAMP Primer Design
RT-LAMP assay has six primers which are two inner primers (FIP and BIP), two outer primers (F3 and B3), and two loop primers (forward loop primer; LF, and backward loop primer; LB) (11, 12). RT-LAMP primers were designed by Primer Explorer (https://primerexplorer.jp/e/) for the N-gene of SARS-CoV-2. For positive control, N-gene sequence was obtained from the National Center of Biotechnology Information (NCBI) with GenBank accession number MN908947.3 and genomic positions between 27894..28259 of Wuhan-Hu-1 genome (NCBI Reference Sequence: NC_045512) was used which was cloned in pGEM®-T easy vector.
RT-LAMP Assay
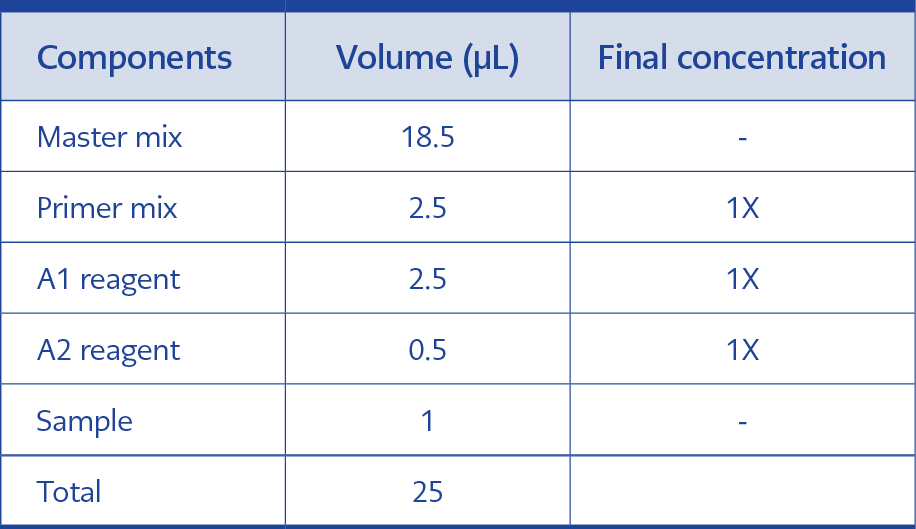
First, the swab samples in virus inactivation medium (vNAT) were lysed with the colorimetric RT-LAMP assay kit lysis buffer at 95 ˚C for 5 min, in which they were mixed at 1:1 portion. Then, the RT-LAMP mix was prepared. For one reaction, the total volume is 25 µL:18.5 µL master mix, 2.5 µL primer mix (10X) (16 µM FIP/BIP, 2 µM F3/B3, 4 µM LF/LB), 2.5 µL A1 (10X), 0.5 µL A2 (50X) and 1 µL was taken from lysed swab sample (Table 1). For positive control, N-gene plasmid (0.5 ng) was mixed with vNAT and quick lysis solution in 2:1:1 proportion. For the negative control, 1:1 portion lysis buffer and vNAT were used. The reaction mixture was incubated at 65 ˚C for 30 min, and at the end of the reaction, the color difference between positive and negative samples was analyzed with the naked eye. The positive samples were seen as yellow in color, and the negative ones were purple.
Testing Clinical RNA Samples with the RT-LAMP Assay
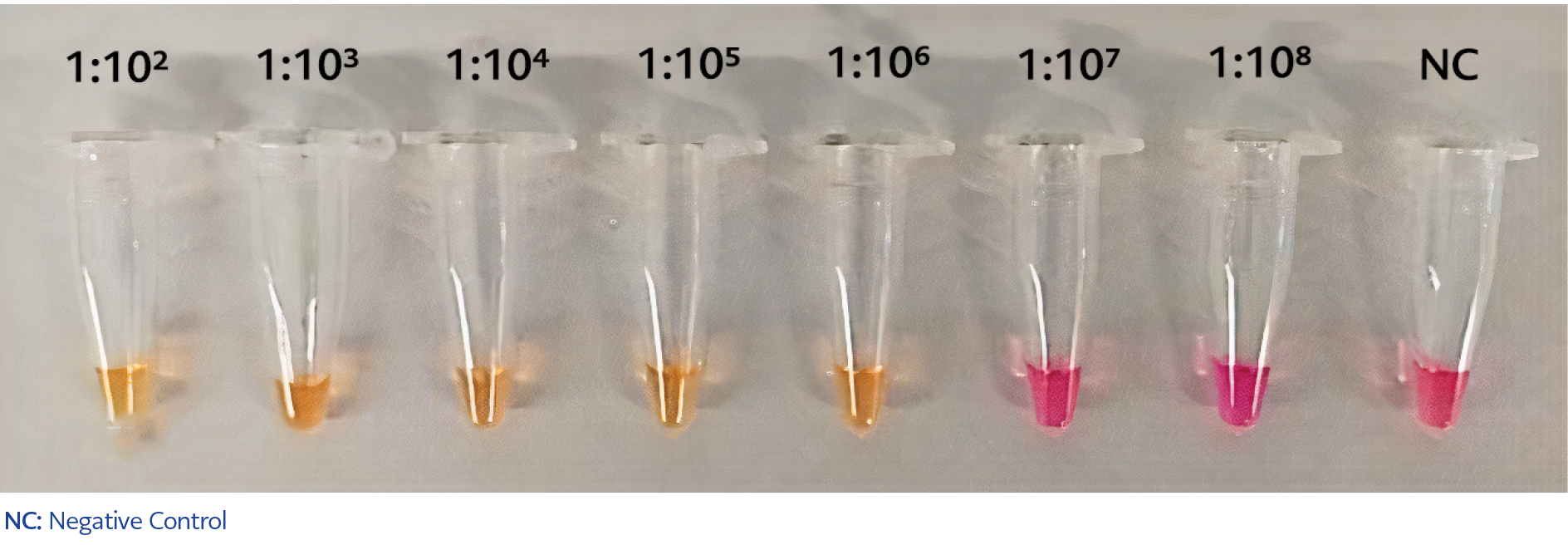

Figure 1. Validating the sensitivity of RT-LAMP assay. The sensitivity of RT-LAMP was determined using 1:102 to 1:108 serial dilutions of the N-gene which was mixed with vNAT and lysis solutions in 2:1:1 proportion.
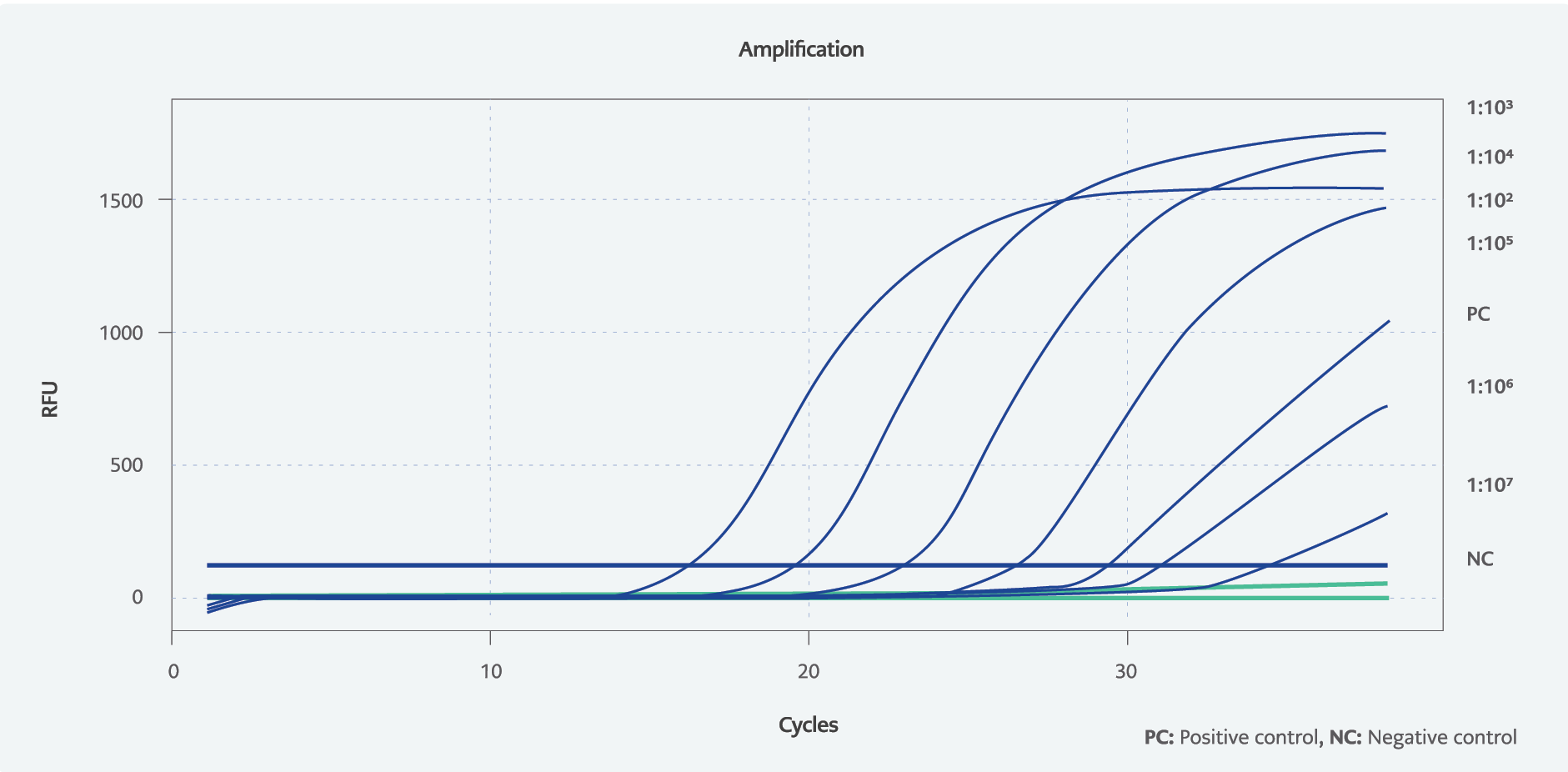
Figure 2. Amplification curves of RT-PCR to detect the sensitivity. Diluted N-gene in vNAT and lysis solution was used to compare the sensitivity with RT-LAMP.
The volunteers of 300 applying for the COVID-19 test at Hacettepe University Hospital from November 3 through November 24, 2020, were included in the study. Of the volunteers, 150 were randomly selected from those who did not show symptoms of COVID-19 and 150 from those who showed symptoms. Combined nose-throat swabs were taken from the volunteers once, and the samples were transferred under appropriate conditions to Hacettepe University Central Laboratory, Department of Molecular Microbiology. The samples were labeled by recording the date, presence of symptoms, and the anonymous number given to the patient. All the samples were tested simultaneously in both the colorimetric RT-LAMP assay kit, we developed to diagnose COVID-19, and the RT-PCR method used in routine practice. Inconsistent test results were further investigated, considering the clinical and epidemiological findings, laboratory results, and radiological evaluations of the volunteers.
Ethics committee approvals (October 13, 2020 dated, 2020-118 numbered subject, 68869993-511.06-E.233430 numbered application) were obtained from the Republic of Turkey Ministry of Health, Turkish Medicines and Medical Devices Agency. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the Helsinki Declaration and its later amendments or comparable ethical standards. Our all database is available to audit. Informed consent was obtained from all individual participants included in the study.
Results
Sensitivity of the RT-LAMP Assay in Detecting SARS-CoV-2 Using N Gene
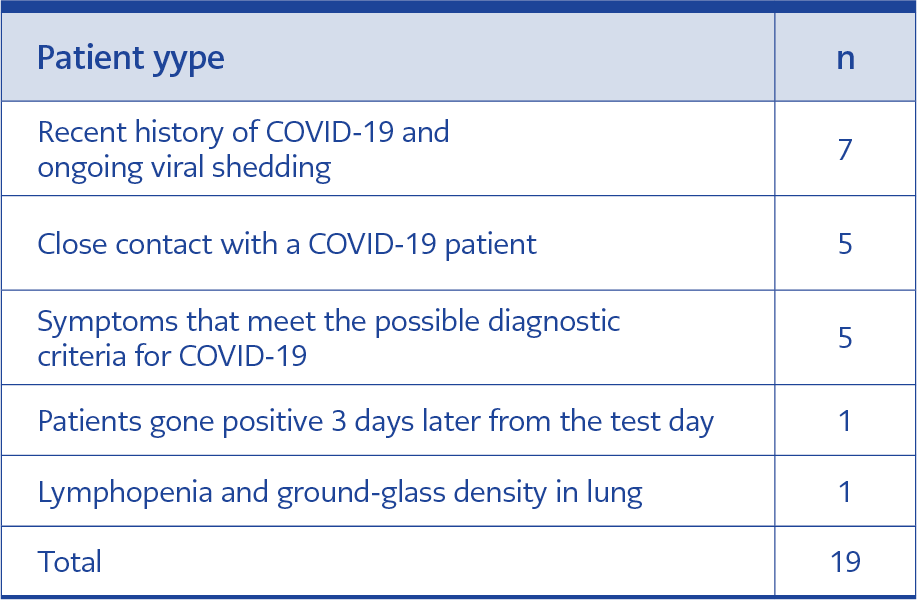

Table 4. Evaluation of volunteers whose clinical, epidemiological, laboratory, and radiological findings supported RT-LAMP assay kit positivity despite negative RT-PCR results.
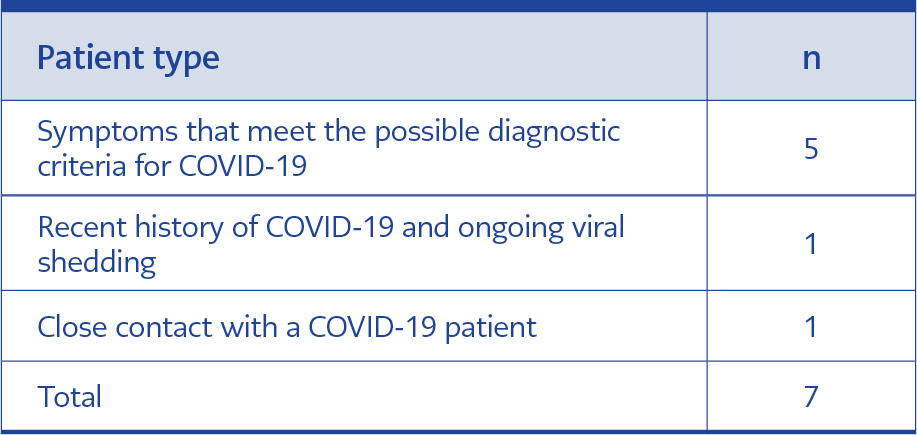

Table 5. Evaluation of volunteers diagnosed with COVID-19 by RT-PCR positivity despite negative results with RT-LAMP assay kit.
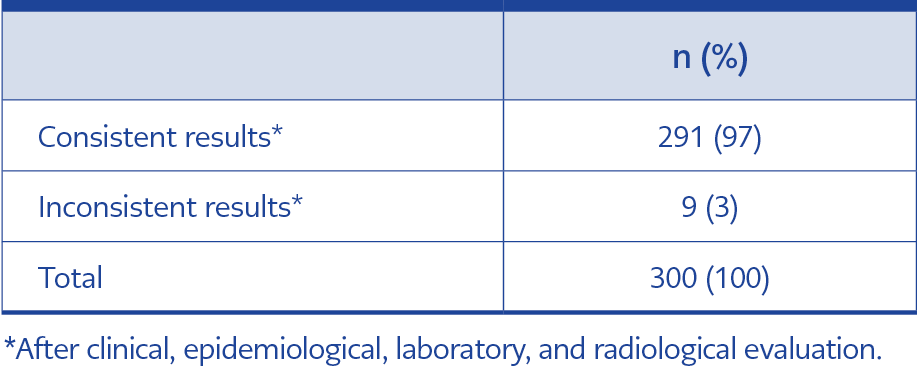
Table 6. Consistency of reassessed results after clinical, epidemiological, laboratory, and radiological evaluations.
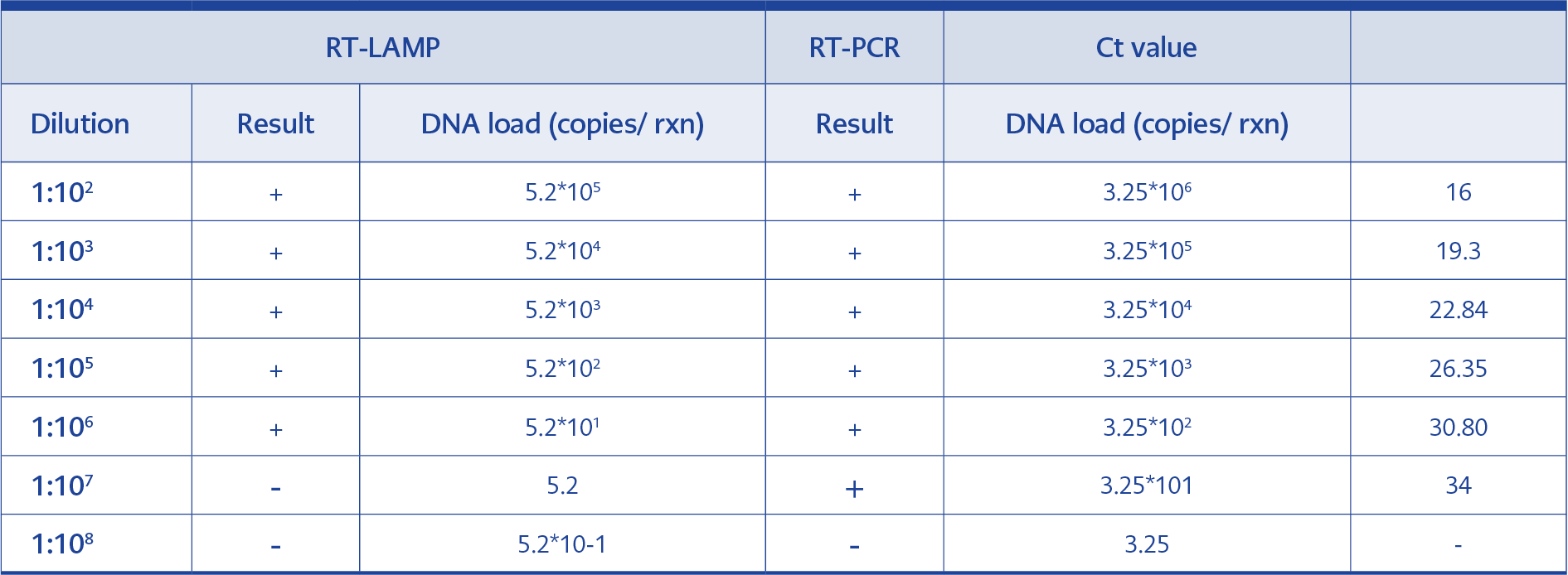
The colorimetric RT-LAMP assay was developed to detect the nucleocapsid (N) gene of SARS-CoV-2. To evaluate the sensitivity of the RT-LAMP assay, serial dilutions of 5*10–3 ng DNA from 1:102 to 1:109 were prepared from the N-gene plasmid. Because of the preparation of the N-gene plasmid in 2:1:1 proportion with vNAT and lysis buffer, the final concentrations of the diluted N-gene plasmid solutions became 5*10-4 to 5*10-9 ng/µL. As a result, the color change from purple to yellow in 30 minutes was observed up to 106 dilutions, equal to 5*10-6 ng (Figure 1). In addition, number of copies of the cloned N-gene was quantified as the stated equation: Copies/μL = concentration of plasmid (g/μL) / [(plasmid length × 660) × (6.022 × 1023)] and our RT-LAMP assay was detected the N-gene up to nearly 60 copies/µL (Table 2) (10).
To validate these results, RT-PCR was performed with these dilutions (Figure 2). For RT-PCR, serial dilutions of 5*10-3 ng/ µL. DNA from 1:102 to 1:109 were prepared from the N-gene plasmid, the same as RT-LAMP. N-gene plasmid was prepared in 1:1 proportion with vNAT, and the final concentrations of the diluted plasmid solution became 1.25*10-2 ng/µL. It was observed that RT-PCR was able to detect approximately 30 copies which means nearly the same as RT-LAMP (Table 2). However, the starting volume of RT-LAMP is 25 µl, and 1µl belongs to the sample. In RT-PCR, the total volume is 20 µl, and 5 µl belongs to the sample. Thus, the total volume is lower, and the sample volume is higher in RT-PCR, in which the copy numbers were calculated according to sample volumes per reaction. The starting DNA volume of RT-PCR is higher and more concentrated than RT-LAMP, so this result can be expected to observe.
Comparison of the RT-LAMP Assay and RT-PCR Results with Clinical Samples
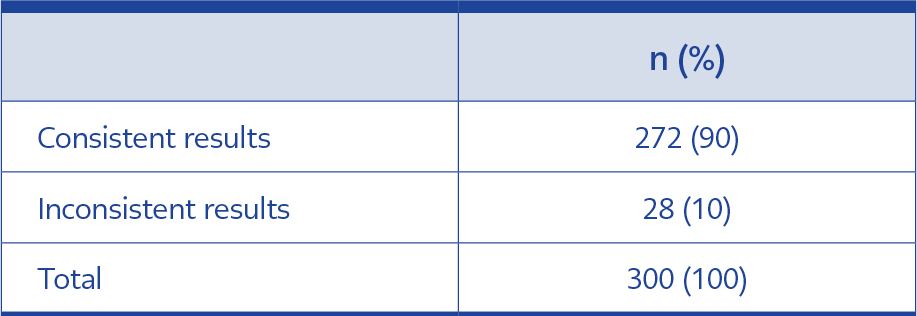
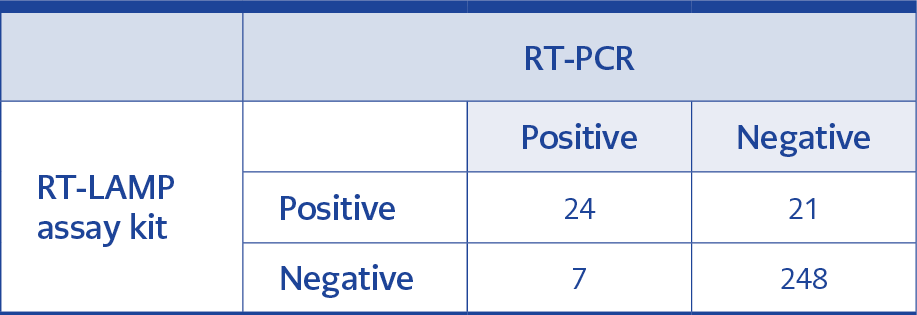
Analysis of the RT-LAMP and RT-PCR test results confirmed 90% consistency; 272 results out of 300 were consistent in both RT-LAMP and RT-PCR (Table 3a). In the inconsistent group, seven samples tested negative with the RT-LAMP assay kit, but these samples belonged to volunteers who were diagnosed with COVID-19 with positive RT-PCR results. The remaining 21 inconsistent samples were positive in the RT-LAMP assay kit, but RT-PCR results were negative (Table 3b).
Then, the inconsistent results were investigated more extensively by other evaluations. Twenty-one samples, which were negative in RT-PCR, were found positive with the RT-LAMP kit. The clinical, epidemiological, laboratory, and radiological evaluations supported RT-LAMP kit positivity for 19 out of 21 patients. Seven of these patients had had COVID-19 recently, but none of them showed symptoms at the time of sample collection, so we concluded that the viral shedding of these patients was continuing. Five of them had close contact with a COVID-19 patient. Five patients had the symptoms, which were possible diagnostic criteria for COVID-19. One patient tested positive for both RT-PCR and RT-LAMP after three days of the first sample collection. Thus, RT-LAMP can detect COVID-19 in its preliminary phases. In the last case, it was determined that one patient had COVID-19 after examining his/her radiology result, and it was diagnosed that the patient had lymphopenia and ground glass density in the lung compatible with early-stage COVID-19. The positivity of the remaining two RT-LAMP test kit results could not be supported by clinical, epidemiological, laboratory, and radiological findings (Table 4).
While the RT-LAMP test kit results of seven volunteers were negative, they tested positive for RT-PCR. Two of these samples were taken from asymptomatic individuals. One of these two individuals had had medium-risk contact with a person diagnosed with COVID-19. The other had had COVID-19 recently, and it was concluded that viral shedding may continue during the convalescence period. The other five samples were taken from patients who already showed symptoms, and the diagnoses of COVID-19 were confirmed by repeated RT-PCR results, which did not support the results of the RT-LAMP test kit (Table 5).
As shown in Table 3a, the RT-LAMP test results of 272 volunteers were consistent, and 28 were inconsistent with RT-PCR. In this case, the consistency between RT-LAMP and RT-PCR was 90%. Being depicted in Table 4, the evaluations of the volunteers for 19 of the 28 inconsistent samples supported the results of RT-LAMP. Based on these data, only nine out of 300 volunteers’ clinical, epidemiological, laboratory, and radiological evaluations were inconsistent between RT-LAMP and RT-PCR. The final evaluation showed a consistency of 97% between RT-LAMP and RT-PCR results (Table 6). Moreover, it was calculated that the colorimetric RT-LAMP assay kit has 98% specificity and 96% sensitivity according to these evaluations. Thus, the RT-LAMP assay kit was considered a strong tool for diagnosing COVID-19 with high sensitivity and specificity.
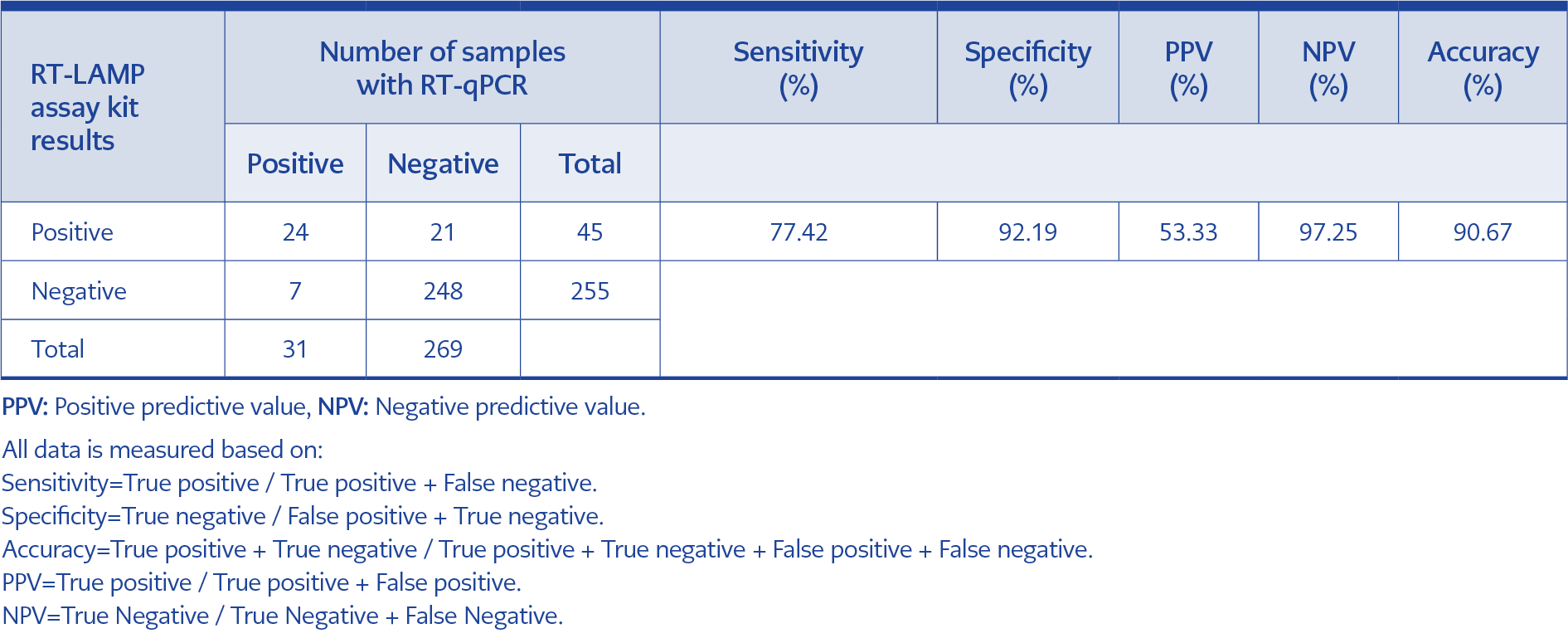
The results of 300 samples tested in RT-LAMP and RT-PCR are demonstrated in Table 6. In all RT-LAMP assay, the reaction tubes illuminated with phenol red was evaluated with the naked eye. Positivity rates of RT-PCR and RT-LAMP were 10.33% (31/300) and 15% (45/300), respectively. The sensitivity of RT-LAMP was 77.42%, and its specificity was 92.19%. The results of the sensitivity, negative predictive value (NPV), positive predictive value (PPV), accuracy, and specificity of RT-LAMP compared with RT-PCR in each group are demonstrated in Table 3c.
Discussion
In this study, we developed a colorimetric RT-LAMP assay for SARS-CoV-2 viral RNA and compared it with RT-PCR. N-gene plasmid serial dilution was performed to test the colorimetric RT-LAMP assay’s sensitivity, and it was significantly comparable with RT-PCR. Reevaluation and further investigation of the inconsistent results demonstrated that 19 out of previously reported positive 21 cases with the RT-LAMP assay kit were truly positive, greatly supporting the specificity and efficiency of the RT-LAMP assay kit. Examining the inconsistencies between RT-PCR and RT-LAMP continued with diagnostic methods like anamnesis, screening, and examining radiology results. Moreover, inconsistent results between RT-PCR and RT-LAMP were investigated with further diagnostic methods like anamnesis, screening, and examining radiology results. According to these findings, 19 of the 21 inconsistent positive results were re-diagnosed as positive.
In line with these results, the Republic of Turkey Ministry of Health, Turkish Medicines and Medical Devices Agency accepted the colorimetric RT-LAMP assay kit as an in vitro diagnostic kit to use detection of COVID-19 in humans with the product tracking system number 8683011276071, even in the early phases of the disease.
The limitations of our study are that it was conducted in a single center, no follow-up visits were made to the patients, and that variant strains were not in circulation when we conducted the study. Another limitation of this study is the lack of plasmid with T7 promoter. Therefore, we did not measure RNA amplified from the plasmid using RT-LAMP and RT-PCR assays.
The strengths of our study are that we studied both symptomatic and asymptomatic individuals when comparing RT-PCR and RT-LAMP results, we were blind to the results because we did not know the RT-PCR results beforehand, and we studied the assays on the same patient sample when making the comparison.
This study shows that the RT-LAMP assay is a good substitute for conventional PCR-based techniques for its rapidity, sensitivity, and unique temperature requirements, thereby providing on-site detection of SARS-CoV-2 without demanding complicated equipment. In addition, RT-LAMP assay results could be evaluated visually. In our study, the colorimetric RT-LAMP assay kit was very sensitive and could detect 52 copies of DNA of SARS-CoV-2. The RT-LAMP assay kit proved to be applicable to SARS-CoV-2 detection, eliminating the need for expensive thermal cyclers, gel electrophoresis, and time-consuming DNA extraction techniques.
Although RT-PCR is the gold standard diagnostic method for COVID-19, rapid and accurate tests are needed to control the pandemic. The colorimetric RT-LAMP assay kit may be a cost-effective, time-saving, and accurate alternative to RT-PCR, the gold standard method.