Origins, characteristics and classification of extended-spectrum beta-lactamases (ESBLs)
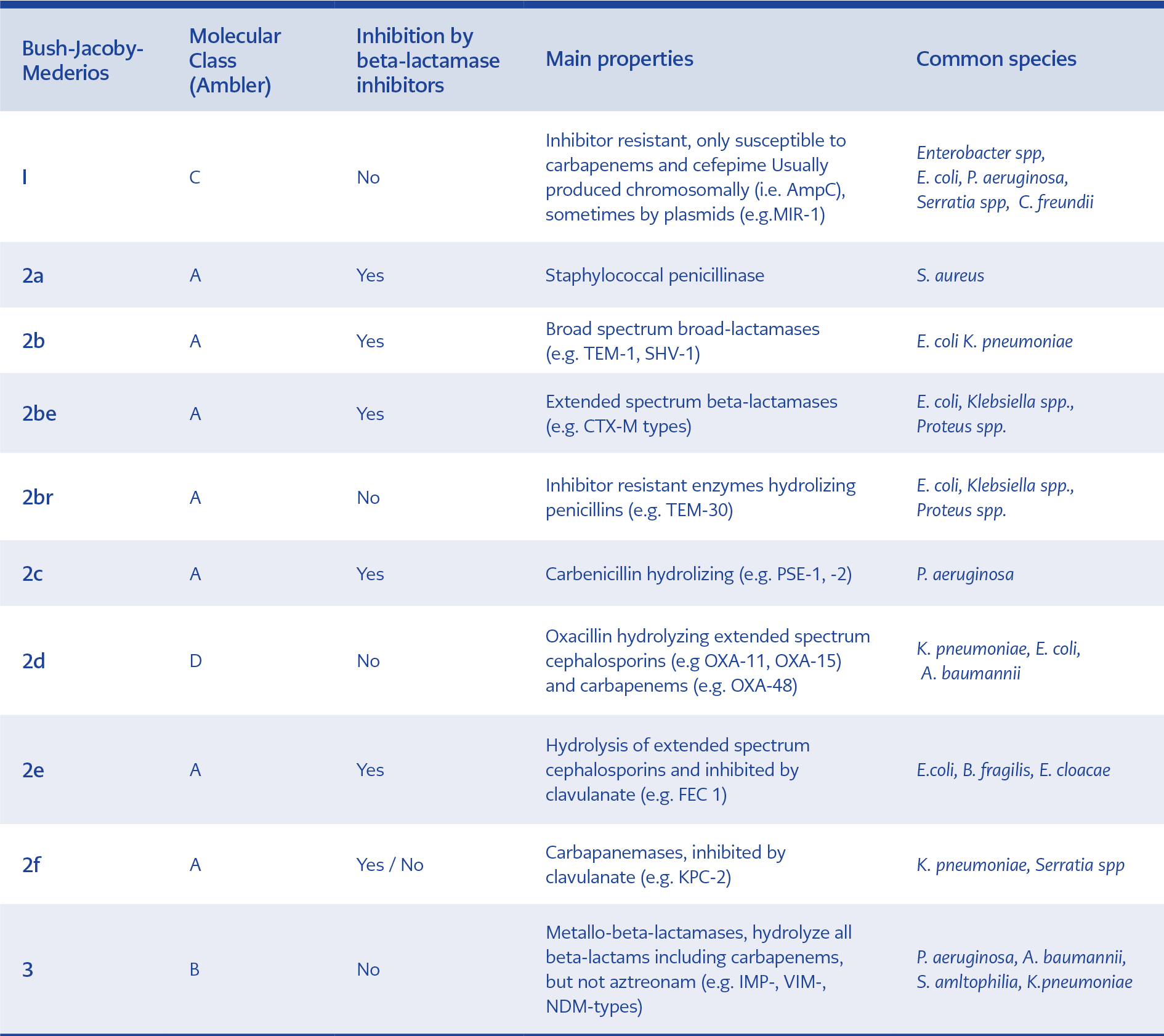
Currently, beta-lactamase family includes more than 2800 unique proteins (1). Although varied types of beta-lactamases have been specified to date, they have common topographic structures consisting of alpha-helices and beta-plated sheets (2). They most probably originated from environmental sources and produced against naturally occurring beta-lactams. Although many classification systems have been proposed for beta-lactamases, the most commonly used ones are Ambler scheme and Bush-Medeiros-Jacoby system according to their amino acid homology and functional properties (substrate and inhibitor profiles), respectively (3,4). Both classification systems are illustrated and compared in Table 1. ESBLs include three major families, TEM, SHV, and CTX-M, with a large variety of other groups of enzymes. The first plasmid-mediated beta-lactamase was isolated in the early 1960s and named as TEM-1 with hydrolytic activity mainly against penicillins and first-generation cephalosporins. In the early 1980s, broad spectrum-cephalosporins (BSCs) were introduced into clinical practice. They were primarily used to treat infections with TEM-1- and SHV-1 penicillinase-producing organisms. The first ESBL-producing K. pneumoniae capable of hydrolysing BSCs was discovered in Germany in 1983. These enzymes mainly carried on plasmids and conferred resistance to penicillins, first, second and third-generation cephalosporins (3rdGCs) and aztreonam. However, carbapenems and cephamycins (e.g. cefoxitin) were stable (5,6). Over the years, ESBL enzymes have evolved via point mutations around the active site of TEM-1 or SHV-1 and gained the ability to hydrolyse different types of beta-lactam antibiotics (7). These mutations broadened the spectrum of activity of new ESBL derivatives by increasing affinity (lowering Km values) of these enzymes against a wide range of beta-lactam antibiotics (8). On the other hand, CTX-M (active on CefoTaXime, first isolated in Munich) type of ESBLs emerged through the mobilisation of chromosomal beta-lactamase (bla) genes from the Kluyvera spp. (9). The first CTX-M was reported from Japan in 1986 and initially named as TOHO-1 and later changed to CTX-M (10). These beta-lactamases hydrolyse cefotaxime and ceftriaxone better than they do ceftazidime. Presently, CTX-M group was classified into five different subgroups according to their amino acid identities and included the CTX-M-1, 2, 8, 9, 25 groups.
ESBLs are usually inhibited by clavulanate and tazobactam. This property discriminates them from AmpC-type beta-lactamases (11). However, insufficient inhibition by tazobactam or clavulanate can be observed in some clinical ESBL-producing isolates. SHV-type ESBLs are generally more resistant to beta-lactamase inhibitors (BLIs) than CTX-M type (12). CTX-M type ESBLs are more efficiently inhibited by tazobactam compared with clavulanate.
Institutions setting standards for in vitro antibiotic susceptibility testing both from Europe and United States (i.e. EUCAST and CLSI) do not recommend routine detection of ESBLs in the microbiology laboratory, claiming ESBL expression is less important than the MIC values in determining the optimal therapy for ESBL-producing Enterobacteriaceae infections. Also, phenotypic detection methods for ESBLs are somewhat cumbersome for microbiology laboratories and require one additional day to obtain results. ESBL detection methods can give false results for pathogens expressing AmpC- or Metallo-beta-lactamases combined with ESBLs and Klebsiella oxytoca which shows ESBL phenotype by a chromosomal enzyme called K 1 that hydrolyses some cephalosporins but not others (13). However, in some cases, reporting the presence of ESBLs may be essential to avoid making significant errors in treatment. Pitout et al. (14) demonstrated that the Vitek-2 microdilution method might fail to detect piperacillin-tazobactam (PTZ) resistance, especially in CTX-M-15 and OXA-1 co-producing isolates. ESBL expression may also compromise clinical outcomes of some antibiotic therapies, although MIC values of these antibiotics are in susceptibility ranges. To that end, reduction of MIC values is not a panacea since phenotypic ESBL detection methods may be more sensitive and specific than CLSI and EUCAST breakpoints to identify the presence of ESBLs. From a clinical standpoint, performing ESBL detection tests routinely in countries having high ESBL prevalence may also be reconsidered for purposes of infection control and epidemiological investigations.
Antibiotic co-resistance mechanisms among ESBL-producing Enterobacteriaceae
The plasmids containing genes expressing ESBLs frequently carry genes conferring resistance to aminoglycosides (AGs), fluoroquinolones (FQs) and trimethoprim-sulfamethoxazole (TMP-SMX) (15, 16). Furthermore, hyperproduction of beta-lactamases, porin mutations, co-production of narrow-spectrum oxacillinases (e.g. OXA-1) and AmpC diminish susceptibility to several antibiotics involving BSCs, beta-lactam/beta-lactamase inhibitor (BLBLI) combinations, and even carbapenems (17). AmpC is usually encoded by chromosomal genes; however, plasmid-mediated AmpC acquisition is not uncommon (18). ESBLs expressed with AmpC may not be reliably detected by routine laboratory methods since >8 fold reduction in MIC values of third-generation cephalosporins cannot be detected with clavulanate when concomitant AmpC expression is present. AmpC-producing pathogens are resistant to BSCs, BLBLIs, and cephamycins, but spare cefepime. The activity of beta-lactam antibiotics can also be substantially impaired by overexpression of parent enzymes (TEM-1 or SHV-1) (19). Furthermore, the simultaneous production of multiple ESBL genes in a given isolate can reduce the effectiveness of BLBLIs (7).
OXA-1/30 (mainly OXA-1) and CTX-M genes can be carried on the same plasmid and may render pathogens resistant against amoxicillin-clavulanate (AMC) and piperacillin-tazobactam (PTZ) (20). OXA 1/30 is frequently associated with CTX-M-15 in both E. coli (particularly in ST131) and K. pneumoniae (21, 22). Additionally, OXA-1 may be carried with aac (6’)-Ib-cr gene which compromises the activities of amikacin and tobramycin. Rapid diagnostic methods revealing clinically significant resistance mechanisms in a short time can prove very useful in such settings.
Clinical and molecular epidemiology of ESBLs
According to the 2017 EARS-Net surveillance; resistance rates to the 3rdGCs were 14.9% and 31.2% in invasive isolates of E. coli and K. penumoniae, respectively, in the European Union (EU). More than 80% of isolates in both species were ESBL producers. The resistance rates were highest in Eastern and Southern European countries as compared with Northern and Western Europe (23). In the US, the incidence of infections with ESBL-producing organisms has increased from 1997 to 2011, slightly more frequent for infections with ESBL-producing Klebsiella spp. compared with ESBL-producing E. coli infections. The ESBL phenotype was identified in 7% of the 2,768 Klebsiella isolates tested in 1997–2000 at 30 US hospitals (24). This figure increased to 15% among isolates collected from 79 US hospitals between 2011 and 2013 (25). For E. coli, two surveillance programs in the US reported a proportion of ESBL phenotype 1% and 8% of isolates in 1997–2000, while in 2011–2013 this proportion raised to 12% (26). In Southeast and East Asia, the nosocomial detection rates of ESBL E. coli were 20–40%, and it has reached 60–70% in China (27). In a recent meta-analysis, the ESBL detection rates in long-term care facilities were reported to be 10–60% in European countries and ~50% in China (28). In another meta-analysis (including 66 studies) concerning with the ESBL faecal colonization prevalence in the community, the pooled prevalence rate was 14% (95% confidence interval, 9.0–20.0%) and the authors predicted that this figure was likely to increase by 5.38% annually (29). The colonization rates were ~4% in Europe, ~2% in North America and ~46% in West Pacific regions.
Molecular epidemiology of ESBLs has changed considerably in the last decade. Currently, blaCTX- M-15 genes are the most prevalent ESBL genes in most regions of the world (30). The blaCTX-M-14 genes are also frequently detected in some parts of Europe, such as Spain, and East and Southeast Asia. Nevertheless, in Southeast Asia, including India, blaCTX- M-15 is more frequent than blaCTX-M-14 (30, 31).
Most of the infections caused by community-onset ESBL-producing Enterobacteriaceae involve urinary tract infections (UTIs). Moreover, some life-threating infections, such as bloodstream infections (BSIs) and intra-abdominal infections (IAIs), have been encountered with increasing frequency (32-34). In the mid to late 1990s, some anecdotal studies reported that ESBL-producing Enterobacteriaceae had started to disseminate from inpatient to outpatient settings (35, 36). One possible reason for this dissemination was the gradual increase of medical care in long-term care facilities (LTCFs) where severely ill patients including those with central lines, urinary catheter, other invasive devices and those are mechanically ventilated were managed for long-term periods (37). Since many of LTCFs are ’for-profit’ organizations, they were initially reluctant to apply infection control and antimicrobial stewardship measures (38). Therefore, LTCFs soon became an essential source of ESBL-producing Enterobacteriaceae, along with other multi-drug resistant organisms (MDROs) (39, 40). Those patients with complicated medical conditions were continually transferred back and forth between health care facilities, and they served as “Trojan horses” for MDROs, including ESBL-producers (41). This evolution of medical care facilitated the spread of formerly pure nosocomial ESBLs into non-hospital settings (32). Recently, Pulcini et al. (42) conducted a large-scale study to identify the differences of antibiotic resistance for microorganisms isolated from urinary samples between community dwellers and nursing home residents. The frequencies of ESBL-producing E. coli were 4.6% and 7.7% (p=<0.0001) in outpatient and inpatient settings, respectively.
Currently, the primary pathogens causing community-onset infections are CTX-M type ESBL-producing E. coli (32). The patients who colonized with these pathogens are often previously treated with FQs and 3rdGCs, are exposed to invasive interventions and nosocomial environments (7). In addition to the selective pressure effect of antibiotic overuse in humans, uncontrolled use of antibiotics in veterinary medicine and food-producing animals has led to the rapid dissemination of ESBLs in the community (43). Additionally, environmental sources such as urban wastewaters, contaminated drinking water, and spreading via international travel have been proposed as possible acquisition means of ESBL-genes in the community (44). However, relative contributions of the factors mentioned above on widespread dissemination of CTX-Ms in the world are still debated.
Horizontal gene transfer is one of the most strong forces in bacterial evolution. When virulent bacterial clones acquire resistance determinant genes, they can emerge as a dominant pathogen through clonal expansion within local or global population such as ST131 as the dominant extra-intestinal pathogenic E. coli worldwide (45). ST131, highly virulent strain of E. coli, has been isolated in various infections including meningitis, osteomyelitis, peritonitis (46, 47), UTIs and urosepsis (48).
Impact of initial inadequate empirical antimicrobial therapy on clinical outcomes
Infections caused by ESBL-producing organisms are associated with a higher rate of mortality (49-51), prolonged infection-related hospital stay (52, 53) and higher healthcare-associated costs (54-56). Therefore, initially selecting appropriate empirical antibiotic therapy is very important, but also a challenge with the rising incidence of antimicrobial resistance. Although some studies asserted no significant impact of inappropriate initial empirical antimicrobial therapy on mortality (57, 58), many other studies consistently demonstrated otherwise (59, 60). This contradiction can be explained by resistance profile of the causative pathogen, source and severity of infections, achieving appropriate source control, baseline comorbidities and place of acquisition of infection (e.g. community-onset vs hospital-acquired). Tumbarello et al. (61) investigated the determinants of inappropriate empiric antibiotic therapy in bacteremia with ESBL-producing E. coli. They concluded that unknown source of bacteremia, resistance to more than three antimicrobials, previous hospitalization and antibiotic exposure were risk factors for receiving inappropriate empiric antibiotic therapy.
Early prediction methods for bloodstream infections with ESBL-producing Enterobacteriaceae and impact of de-escalation on clinical outcomes
The World Health Organization (WHO) published a global priority pathogens list to focus attention on the most hazardous pathogens for public health. Enterobacteriaceae resistant to the 3rdGCs (which includes ESBL-producing Enterobacteriaceae) were included within the critical category (first priority) of this list because of a rapid increase in prevalence particularly in community, the easy transmission of ESBL genes via plasmids and limitations in antibiotic choices for treatment of infections caused by these pathogens. However, timely identification of ESBL-producing bacterial infections can improve relevant outcomes. Incorporation of ESBL-prediction scores may improve the appropriateness of empirical antimicrobial therapy and reduce carbapenem use. To that end, Goodman and colleagues (62) developed a decision tree algorithm to estimate the likelihood of a bacteremic patient being infected with an ESBL-producing E. coli or Klebsiella spp. The final tree that stratifies bacteremia with Enterobacteriaceae according to the risk of ESBL production contained five predictors: the history of prior ESBL colonization and infection, chronic indwelling vascular hardware, age ≥43 years, recent hospitalization in an ESBL high burden region and ³6 days of antibiotic exposure in the prior six months. The positive and negative predictive values of this decision tree were 90.8% and 91.9%, respectively. Sensitivity rate was 51.0%, and specificity rate was 99.1%. In another study, multiple logistic regression analysis was used to identify independent risk factors for BSIs with ESBL-producing Enterobacteriaceae (63). Prior colonization/infection with ESBL-producing Enterobacteriaceae, outpatient gastrointestinal or genitourinary procedures within one month and the number of prior courses of beta-lactams and FQs used within the previous three months were independent risk factors for BSIs with Enterobacteriaceae. However, these decision tree analysis and early prediction risk score models need to be validated in large scale studies.
Early prediction of ESBL-producing organisms as a causative pathogen via prediction scores or machine learning may enable us to give appropriate early therapy (64). After antimicrobial susceptibility results, the initial therapy may be streamlined and narrowed, if possible. Two different research groups investigated the impact of this practice, so-called “de-escalation or step-down therapy” on clinically relevant outcomes in BSIs with Enterobacteriaceae. They found that both oral step-down therapy and early de-escalation of therapy were not associated with higher 30-day mortality rates and clinical failure rates in BSIs with Enterobacteriaceae. Furthermore, switch to oral therapy shortened the duration of hospitalization significantly (65, 66).
Conclusion
ESBL-producing Enterobacteriaceae species are one of the most frequently encountered pathogens in both hospital-acquired and community-onset infections. Their widespread dissemination, particularly in the community, has threatened public health for the last two decades. Several high-risk clones and successful plasmid types have played a significant role, particularly in the dissemination of ESBL-producing E. coli worldwide. The wide array of co-resistance mechanisms has also rendered many different classes of antibiotics useless. Therefore, delayed initiation of appropriate antimicrobial therapy was frequently reported in previous studies concerning infections caused by ESBL-producing Enterobacteriaceae. In order to resolve this issue, rapid diagnostic methods should be explored to shorten interval time between taking a sample for culture and identification of species, in-vitro susceptibilities and resistance determinants. Rapid diagnostics will provide relevant information not only for the determination of appropriate therapy much earlier but also for implementing the appropriate infection control measures and epidemiological investigations. To that end, affordable, user-friendly and accurate rapid diagnostics are urgently needed in routine practice. Also, prediction score models or decision tree algorithms may be useful for early prediction of infections caused by ESBL-producing organisms and prescribing appropriate antimicrobial therapy for infections caused by these pathogens. Similarly, machine learning may be an appealing tool in the near future for this purpose.